Developmental and epileptic encephalopathy (DEE) refers to a group of neurodevelopmental conditions characterized by developmental delay, cognitive impairment, and seizures in children. In 2016, the first case linking variants in both the copies of UBA5 gene to DEE44 was reported. Since then, twelve distinct missense variants in the UBA5 gene have been identified in 25 patients.
A recent study by Dr. Hugo J. Bellen and his team at the Jan and Dan Duncan Neurological Research Institute (Duncan NRI) at Texas Children’s Hospital and Baylor College of Medicine generated a new fruit fly model to assess the severity of symptoms caused by each of these variants. This systematic analysis of UBA5 pathogenic variants lays the foundation for better evaluation of the variants, which is important for the DEE44 patients in the future and for the development of drugs and gene therapy to treat this rare disorder. The study was published recently in the journal eLife.
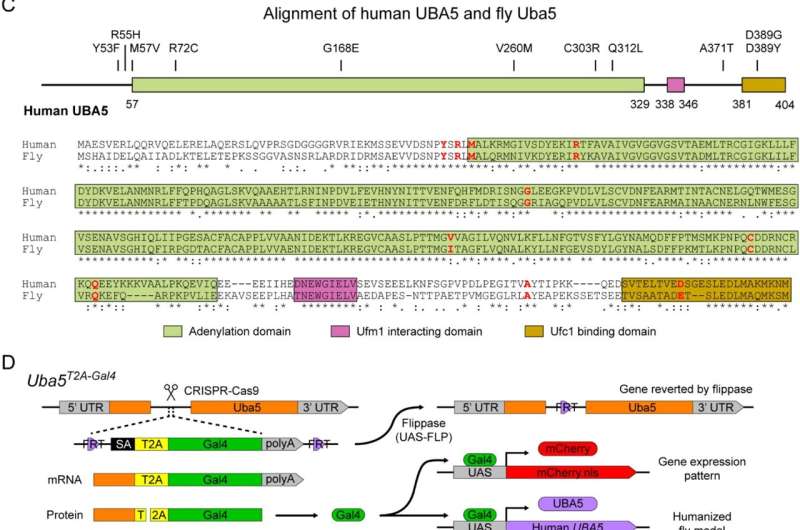
The UBA5 gene is highly conserved across species ranging from fruit flies to humans. It encodes a protein that binds to Ubiquitin-fold Modifier 1 (UFM1) and was discovered two decades ago. It acts as an enzyme that catalyzes a post-translational modification called UFMylation. The exact mechanisms and biological significance of UFMylation are still unclear, but it adds a new regulatory layer to many fundamental cellular processes.
“Since UBA5 variants in patients have been found in precisely the same amino acid residues in fruit fly Uba5 protein, we generated a humanized model by expressing the human version of the UBA5 gene in fruit flies lacking the endogenous Uba5 gene,” said Dr. Xueyang Pan, who is the first author of the study and a postdoctoral fellow in the Bellen lab. “Next, we set out to characterize the strength of the disease-causing variants by expressing them individually in flies and observing the phenotypes caused by the variants.”
They found that the severity and the type of symptoms varied widely among different variants. For instance, they found four variants that failed to rescue the lethality caused by the loss of endogenous Uba5. Another five variants caused progressive motor defects, three of which also caused developmental delays or seizure-like symptoms. While the normal human UBA5 substituted all essential functions of its fly counterpart, the disease-causing variants exhibited variable loss of function that were classified as mild, intermediate, or severe subtypes. Importantly, exogenous overexpression of the UBA5 gene did not cause any obvious defects, suggesting a potentially safe therapeutic strategy to increase UBA5 enzyme levels, which is encouraging.
In collaboration with Dr. Jonathan Pruneda and Dr. Ruth Napier at Oregon Health & Science University and the Raiden Science Foundation of Beaverton, Oregon, the team also established several new biochemical assays to determine the impact of these disease alleles on the stability and activity of the UBA5 enzyme. The team found a close correlation between the severity of the symptoms exhibited by a particular variant in fruit flies and its enzymatic stability and/or function, as measured by biochemical assays. These findings provide a molecular explanation for the loss of function among UBA5 variants, and importantly establish a route to develop new drugs that restore UBA5 function and treat the disease.
“We undertook this study because there was a clear need to assess the disease-associated variants of UBA5,” Dr. Bellen said. “This study not only established a powerful experimental paradigm to better understand the etiology of DEE44 but can also serve as a template to better understand other human disorders and to develop new therapies for DEE44 and others.”
Others involved in the study were Albert N. Alvarez, Mengqi Ma, Shenzhao Lu, Michael W. Crawford, Lauren C. Briere, Oguz Kanca, Shinya Yamamoto, David A. Sweetser, Jenny L. Wilson, Ruth J. Napier, and Jonathan N. Pruneda.
More information:
Xueyang Pan et al, Allelic strengths of encephalopathy-associated UBA5 variants correlate between in vivo and in vitro assays, eLife (2023). DOI: 10.7554/eLife.89891.1
Journal information:
eLife
Source: Read Full Article





